DeepDive Tutorial Extracting mentions of spouses from the news
NOTE: we recommend using the Jupyter Notebook version of this tutorial that can be easily launched inside Docker containers.
In this tutorial, we show an example of a prototypical task that DeepDive is often applied to: extraction of structured information from unstructured or 'dark' data such as web pages, text documents, images, etc. While DeepDive can be used as a more general platform for statistical learning and data processing, most of the tooling described herein has been built for this type of use case, based on our experience of successfully applying DeepDive to a variety of real-world problems of this type.
In this setting, our goal is to take in a set of unstructured (and/or structured) inputs, and populate a relational database table with extracted outputs, along with marginal probabilities for each extraction representing DeepDive's confidence in the extraction. More formally, we write a DeepDive application to extract mentions of relations and their constituent entities or attributes, according to a specified schema; this task is often referred to as relation extraction.* Accordingly, we'll walk through an example scenario where we wish to extract mentions of two people being spouses from news articles.
The high-level steps we'll follow are:
Data processing. First, we'll load the raw corpus, add NLP markups, extract a set of candidate relation mentions, and a sparse feature representation of each.
Distant supervision with data and rules. Next, we'll use various strategies to provide supervision for our dataset, so that we can use machine learning to learn the weights of a model.
Learning and inference: model specification. Then, we'll specify the high-level configuration of our model.
Error analysis and debugging. Finally, we'll show how to use DeepDive's labeling, error analysis and debugging tools.
*Note the distinction between extraction of true, i.e., factual, relations and extraction of mentions of relations. In this tutorial, we do the latter, however DeepDive supports further downstream methods for tackling the former task in a principled manner.
Whenever something isn't clear, you can always refer to the complete example code at examples/spouse/ that contains everything shown in this document.
0. Preparation
First of all, make sure that DeepDive has been installed.
Next, DeepDive will store all data—input, intermediate, output, etc.—in a relational database.
Currently, Postgres, Greenplum, and MySQL are supported; however, Greenplum or Postgres are strongly recommended.
To set the location of this database, we need to configure a URL in the db.url file, e.g.:
echo "postgresql://$USER@$HOSTNAME:5432/deepdive_spouse_$USER" >db.url
Note: DeepDive will drop and then create this database if run from scratch—beware of pointing to an existing populated one!
1. Data processing
In this section, we'll generate the traditional inputs of a statistical learning-type problem: candidate spouse relations, represented by a set of features, which we will aim to classify as actual relation mentions or not.
We'll do this in four basic steps:
- Loading raw input data
- Adding NLP markups
- Extracting candidate relation mentions
- Extracting features for each candidate
1.1. Loading raw input data
Our first task is to download and load the raw text of a corpus of news articles provided by Signal Media into an articles table in our database.
We create a simple shell script that downloads and outputs the news articles in TSV format.
DeepDive will automatically create the table, execute the script and load the table if we save it as:
input/articles.tsv.sh
The aforementioned script reads a sample of the corpus (provided as lines of JSON objects), and then using the jq language extracts the fields id (for document id) and content from each entry and converts those to TSV format.
Next, we need to declare the schema of this articles table in our app.ddlog file; we add the following lines:
articles(
id text,
content text
).
Then we compile our application, as we must do whenever we change app.ddlog:
deepdive compile
Finally, we tell DeepDive to execute the steps to load the articles table using the input/articles.tsv.sh script. You must have the full corpus downloaded.
deepdive do articles
Alternatively, a sample of 1000 articles can be loaded directly by issuing the following command:
bash
deepdive load articles input/articles-1000.tsv.bz2
DeepDive will output an execution plan, which will pop up in your default text editor. Save and exit to accept. DeepDive will run, creating the table and then fetching and loading the data.
After that finishes, we can take a look at the loaded data using the following deepdive query command, which enumerates the values for the id column of the articles table:
deepdive query '?- articles(id, _).'
id
--------------------------------------
340bb625-bb7e-49af-aa8d-781e5762f7a3
fe4d045d-6c97-4ccf-a7d5-08aafb72a0db
3585cd84-abd9-4f14-90ff-4fc23b45c84f
eb571ae7-cd9f-4ad2-90a1-a74854574dc9
6354c1a9-8f85-43c6-98d8-6b559d6727ef
8994a0f1-7ca5-45a7-88c6-35bb98e89c95
8b7fe0b3-ef78-4555-bb7a-5ad7007f9a93
74ae2f47-ed24-4768-b8db-980f128061da
89d7ee41-1b11-4f90-b1c9-e69c0145227d
8f9d6a38-a828-40c5-b823-13447e96b842
71984e17-d556-4828-93d3-18daf55a677f
ce117e2b-73cb-48bc-9896-f9c9cb232a7e
06b83cb4-1d03-42e9-8429-6cd272cd4a89
f3397ce9-ba4f-4ddf-93c5-42333b98b907
66ffaca4-4936-4844-a58e-b1a0b02c8df1
84b24e59-a2ee-4eb4-a571-916d83ee475b
3fdcbae1-124e-46af-9773-2401683d19fd
e94a28ec-d335-46c5-aa7f-f14dfdc7258a
69a48149-d459-404f-adbe-6cd6049f71ee
ac5fbfd9-3f62-44bf-ab5a-36ffcd77adb1
70f5365b-51be-40f9-8fe4-380b6010e767
3f328bf8-45a3-441a-9426-2206a8b60b21
55021a16-6d82-4dcf-acef-21d1497910b1
fcd8d601-cce7-4735-92c0-eae086404d44
3a6277d7-8d1f-43bc-a69e-53f640a651a1
e1d7be87-2272-4091-89cb-173939b796a7
48eb9031-30b8-4689-987a-5ee3289448ee
84e7269f-3455-4af6-a1d4-541ae5b34e79
[...]
1.2. Adding NLP markups
Next, we'll use Stanford's CoreNLP natural language processing (NLP) system to add useful markups and structure to our input data.
This step will split up our articles into sentences and their component tokens (roughly, the words).
Additionally, we'll get lemmas (normalized word forms), part-of-speech (POS) tags, named entity recognition (NER) tags, and a dependency parse of the sentence.
We declare the output schema of this step in app.ddlog:
sentences(
doc_id text,
sentence_index int,
sentence_text text,
tokens text[],
lemmas text[],
pos_tags text[],
ner_tags text[],
doc_offsets int[],
dep_types text[],
dep_tokens int[]
).
Next we declare a DDlog function which takes in the doc_id and content for an article and returns rows conforming to the sentences schema we just declared, using the user-defined function (UDF) in udf/nlp_markup.sh.
This UDF is a Bash script which calls our own wrapper around CoreNLP. The CoreNLP library requires Java 8 to run.
function nlp_markup over (
doc_id text,
content text
) returns rows like sentences
implementation "udf/nlp_markup.sh" handles tsv lines.
Finally, we specify that this nlp_markup function should be run over each row from articles, and the output appended to sentences:
sentences += nlp_markup(doc_id, content) :-
articles(doc_id, content).
Again, to execute, we compile and then run:
deepdive compile
deepdive do sentences
Now, if we take a look at a sample of the NLP markups, they will have tokens and NER tags that look like the following:
deepdive query '
doc_id, index, tokens, ner_tags | 5
?- sentences(doc_id, index, text, tokens, lemmas, pos_tags, ner_tags, _, _, _).
'
doc_id | index | tokens | ner_tags
--------------------------------------+-------+-------------------------------------------------------------------------------------
-------------------------------------------------------------------------------------------------------+----------------------------
---------------------------------------------------------------------------------------
36dd2baa-3ce4-4cfa-aac8-6226bb727bb9 | 1 | {Private,automobiles,are,prohibitively,expensive,in,Bethel,",",Alaska,",",and,so,is,
gas,.} | {O,O,O,O,O,O,LOCATION,O,LOC
ATION,O,O,O,O,O,O}
36dd2baa-3ce4-4cfa-aac8-6226bb727bb9 | 2 | {Public,transit,is,nonexistent,.}
| {O,O,O,O,O}
36dd2baa-3ce4-4cfa-aac8-6226bb727bb9 | 3 | {BETHEL,",",Alaska,--,Cheri,Boisvert,and,Matt,Janz,are,moving,apartments,on,foot,in,
the,biggest,town,in,Alaska,'s,Yukon,Kuskokwim,De,.} | {LOCATION,O,LOCATION,O,PERS
ON,PERSON,O,PERSON,PERSON,O,O,O,O,O,O,O,O,O,O,LOCATION,O,LOCATION,LOCATION,LOCATION,O}
36dd2baa-3ce4-4cfa-aac8-6226bb727bb9 | 4 | {``,Bethel,'s,a,strange,place,",",'',Boisvert,said,",",when,I,expressed,surprise,tha
t,she,would,attempt,a,move,by,walking,her,possessions,across,the,dusty,blocks,between,her,two,homes,.} | {O,LOCATION,O,O,O,O,O,O,PER
SON,O,O,O,O,O,O,O,O,O,O,O,O,O,O,O,O,O,O,O,O,O,O,NUMBER,O,O}
36dd2baa-3ce4-4cfa-aac8-6226bb727bb9 | 5 | {But,like,just,about,everybody,else,in,this,city,of,"6,000",",",they,do,n't,have,a,c
ar,",",so,Boisvert,",",a,librarian,",",walks,with,a,tr,can,.} | {O,O,O,O,O,O,O,O,O,O,NUMBER
,O,O,O,O,O,O,O,O,O,PERSON,O,O,O,O,O,O,O,O,O,O}
(5 rows)
Note that the previous steps—here, loading the articles—will not be re-run unless we specify that they should be, using, e.g.:
deepdive mark todo articles
1.3. Extracting candidate relation mentions
Mentions of people
Once again we first declare the schema:
person_mention(
mention_id text,
mention_text text,
doc_id text,
sentence_index int,
begin_index int,
end_index int
).
We will be storing each person as a row referencing a sentence with beginning and ending indexes. Again, we next declare a function that references a UDF and takes as input the sentence tokens and NER tags:
function map_person_mention over (
doc_id text,
sentence_index int,
tokens text[],
ner_tags text[]
) returns rows like person_mention
implementation "udf/map_person_mention.py" handles tsv lines.
We'll write a simple UDF in Python that will tag spans of contiguous tokens with the NER tag PERSON as person mentions (i.e., we'll essentially rely on CoreNLP's NER module).
Note that we've already used a Bash script as a UDF, and indeed any programming language can be used.
(DeepDive will just check the path specified in the top line, e.g., #!/usr/bin/env python.)
However, DeepDive provides some convenient utilities for Python UDFs which handle all IO encoding/decoding.
To write our UDF (udf/map_person_mention.py), we'll start by specifying that our UDF will handle TSV lines (as specified in the DDlog above).
Additionally, we'll specify the exact type schema of both input and output, which DeepDive will check for us:
#!/usr/bin/env python
from deepdive import *
# for python 3 compatibility
try:
xrange
except NameError:
xrange = range
@tsj_extractor
@returns(lambda
mention_id = "text",
mention_text = "text",
doc_id = "text",
sentence_index = "int",
begin_index = "int",
end_index = "int",
:[])
def extract(
doc_id = "text",
sentence_index = "int",
tokens = "text[]",
ner_tags = "text[]",
):
"""
Finds phrases that are continuous words tagged with PERSON.
"""
num_tokens = len(ner_tags)
# find all first indexes of series of tokens tagged as PERSON
first_indexes = (i for i in xrange(num_tokens) if ner_tags[i] == "PERSON" and (i == 0 or ner_tags[i-1] != "PERSON"))
for begin_index in first_indexes:
# find the end of the PERSON phrase (consecutive tokens tagged as PERSON)
end_index = begin_index + 1
while end_index < num_tokens and ner_tags[end_index] == "PERSON":
end_index += 1
end_index -= 1
# generate a mention identifier
mention_id = "%s_%d_%d_%d" % (doc_id, sentence_index, begin_index, end_index)
mention_text = " ".join(map(lambda i: tokens[i], xrange(begin_index, end_index + 1)))
# Output a tuple for each PERSON phrase
yield [
mention_id,
mention_text,
doc_id,
sentence_index,
begin_index,
end_index,
]
Above, we write a simple function which extracts and tags all subsequences of tokens having the NER tag "PERSON".
Note that the extract function must be a generator (i.e., use a yield statement to return output rows).
Finally, we specify that the function will be applied to rows from the sentences table and append to the person_mention table:
person_mention += map_person_mention(
doc_id, sentence_index, tokens, ner_tags
) :-
sentences(doc_id, sentence_index, _, tokens, _, _, ner_tags, _, _, _).
Again, to run, just compile and execute as in previous steps:
deepdive compile && deepdive do person_mention
Now, the person_mention table should hold rows that look like the following:
deepdive query '
name, doc, sentence, begin, end | 20
?- person_mention(p_id, name, doc, sentence, begin, end).
'
name | doc | sentence | begin | end
-------------------+--------------------------------------+----------+-------+-----
Juliette Barnes | d9b82bc6-efa3-4c10-b595-8c51bbe27f3c | 1 | 5 | 6
Hayden Panettiere | d9b82bc6-efa3-4c10-b595-8c51bbe27f3c | 1 | 8 | 9
Shkreli | 85fb02bf-6105-4dfb-995b-bd94a15a9e6d | 10 | 4 | 4
Benjamin Davies | 85fb02bf-6105-4dfb-995b-bd94a15a9e6d | 11 | 5 | 6
Shkreli | 85fb02bf-6105-4dfb-995b-bd94a15a9e6d | 12 | 9 | 9
Alan Craze | ddf3dfd2-cc21-46ca-a0e7-bd66eeb6bec6 | 2 | 3 | 4
Mary Shipstone | ddf3dfd2-cc21-46ca-a0e7-bd66eeb6bec6 | 4 | 0 | 1
Maryam Alromisse | ddf3dfd2-cc21-46ca-a0e7-bd66eeb6bec6 | 4 | 6 | 7
Yasser Alromisse | ddf3dfd2-cc21-46ca-a0e7-bd66eeb6bec6 | 5 | 3 | 4
Mary | ddf3dfd2-cc21-46ca-a0e7-bd66eeb6bec6 | 6 | 9 | 9
Lyndsey Shipstone | ddf3dfd2-cc21-46ca-a0e7-bd66eeb6bec6 | 6 | 15 | 16
Alromisse | ddf3dfd2-cc21-46ca-a0e7-bd66eeb6bec6 | 7 | 7 | 7
Alromisse | ddf3dfd2-cc21-46ca-a0e7-bd66eeb6bec6 | 8 | 0 | 0
Craze | ddf3dfd2-cc21-46ca-a0e7-bd66eeb6bec6 | 10 | 1 | 1
Mary | ddf3dfd2-cc21-46ca-a0e7-bd66eeb6bec6 | 10 | 4 | 4
Alromisse | ddf3dfd2-cc21-46ca-a0e7-bd66eeb6bec6 | 11 | 4 | 4
Chris Isaak | d9b2b908-190f-4010-a5b8-9a083f2b7fe5 | 15 | 0 | 1
Guy Sebastian | d9b2b908-190f-4010-a5b8-9a083f2b7fe5 | 15 | 16 | 17
Dannii Minogue | d9b2b908-190f-4010-a5b8-9a083f2b7fe5 | 15 | 19 | 20
James Blunt | d9b2b908-190f-4010-a5b8-9a083f2b7fe5 | 15 | 22 | 23
(20 rows)
Mentions of spouses (pairs of people)
Next, we'll take all pairs of non-overlapping person mentions that co-occur in a sentence with less than 5 people total, and consider these as the set of potential ('candidate') spouse mentions.
We thus filter out sentences with large numbers of people for the purposes of this tutorial; however, these could be included if desired.
Again, to start, we declare the schema for our spouse_candidate table—here just the two names, and the two person_mention IDs referred to:
spouse_candidate(
p1_id text,
p1_name text,
p2_id text,
p2_name text
).
Next, for this operation we don't use any UDF script, instead relying entirely on DDlog operations. We simply construct a table of person counts, and then do a join with our filtering conditions. In DDlog this looks like:
num_people(doc_id, sentence_index, COUNT(p)) :-
person_mention(p, _, doc_id, sentence_index, _, _).
spouse_candidate(p1, p1_name, p2, p2_name) :-
num_people(same_doc, same_sentence, num_p),
person_mention(p1, p1_name, same_doc, same_sentence, p1_begin, _),
person_mention(p2, p2_name, same_doc, same_sentence, p2_begin, _),
num_p < 5,
p1_name != p2_name,
p1_begin != p2_begin.
Again, to run, just compile and execute as in previous steps.
deepdive compile && deepdive do spouse_candidate
Now, the rows in the spouse candidates should look like the following:
deepdive query '
name1, name2, doc, sentence | 20
?- spouse_candidate(p1, name1, p2, name2),
person_mention(p1, _, doc, sentence, _, _).
'
name1 | name2 | doc | sentence
--------------------+-----------------------+--------------------------------------+----------
William Larnach | Eliza | ab9b3aa5-1b50-4dca-a179-fc2da32cd53e | 40
Jonathan Kraft | Richard M. Berman | c94ecb16-3f32-42b3-8982-19c3ce8c0177 | 11
Jonathan Kraft | Brady | c94ecb16-3f32-42b3-8982-19c3ce8c0177 | 11
Jonathan Kraft | Roger Goodell | c94ecb16-3f32-42b3-8982-19c3ce8c0177 | 11
Nirmala Sitharaman | Biden Mumbai News.Net | e505b558-5db9-4d92-aec0-3b32341499b8 | 8
Nirmala Sitharaman | Joe Biden | e505b558-5db9-4d92-aec0-3b32341499b8 | 8
Nirmala Sitharaman | Husain Haqqani | e505b558-5db9-4d92-aec0-3b32341499b8 | 8
Cecil | Palmer | 118e3e65-9264-4c3d-b9b1-c98a3aed3535 | 3
Mason | Mason | 89ad2fcc-9cb5-41f1-9bd3-0123fcef3ac9 | 38
Palmer | Cecil | 118e3e65-9264-4c3d-b9b1-c98a3aed3535 | 9
Cecil | Palmer | 118e3e65-9264-4c3d-b9b1-c98a3aed3535 | 9
Barker | Constance | ab9b3aa5-1b50-4dca-a179-fc2da32cd53e | 18
Barker | Jill Moon | ab9b3aa5-1b50-4dca-a179-fc2da32cd53e | 18
Steph Laberis | Marty | 8b39759d-835b-4b69-84a4-8359453be9cb | 91
Steph Laberis | Marty | 8b39759d-835b-4b69-84a4-8359453be9cb | 91
Heyde | Theodor Eicke | 1b48ada9-ae5e-4315-b032-95539bdb31d2 | 9
Rustam Kupaisinov | Krister Petersson | 7d429da5-d18a-41ef-a416-9c15b088952e | 2
Rustam Kupaisinov | Yury Zhukovsky | 7d429da5-d18a-41ef-a416-9c15b088952e | 2
Jimmy Tarbuck | Coleen Nolan | 8bb7e4ae-828e-417a-8a15-b182a6bab1e1 | 4
Jimmy Tarbuck | Tarbuck | 8bb7e4ae-828e-417a-8a15-b182a6bab1e1 | 4
(20 rows)
1.4. Extracting features for each candidate
Finally, we will extract a set of features for each candidate:
spouse_feature(
p1_id text,
p2_id text,
feature text
).
The goal here is to represent each spouse candidate mention by a set of attributes or features which capture at least the key aspects of the mention, and then let a machine learning model learn how much each feature is correlated with our decision variable ('is this a spouse mention?').
For those who have worked with machine learning systems before, note that we are using a sparse storage representation-
you could think of a spouse candidate (p1_id, p2_id) as being represented by a vector of length L = COUNT(DISTINCT feature), consisting of all zeros except for at the indexes specified by the rows with key (p1_id, p2_id).
DeepDive includes an automatic feature generation library, DDlib, which we will use here.
Although many state-of-the-art applications have been built using purely DDlib-generated features, others can be used and/or added as well.
To use DDlib, we create a list of ddlib.Word objects, two ddlib.Span objects, and then use the function get_generic_features_relation, as shown in the following Python code for udf/extract_spouse_features.py:
#!/usr/bin/env python
from deepdive import *
import ddlib
@tsj_extractor
@returns(lambda
p1_id = "text",
p2_id = "text",
feature = "text",
:[])
def extract(
p1_id = "text",
p2_id = "text",
p1_begin_index = "int",
p1_end_index = "int",
p2_begin_index = "int",
p2_end_index = "int",
doc_id = "text",
sent_index = "int",
tokens = "text[]",
lemmas = "text[]",
pos_tags = "text[]",
ner_tags = "text[]",
dep_types = "text[]",
dep_parents = "int[]",
):
"""
Uses DDLIB to generate features for the spouse relation.
"""
# Create a DDLIB sentence object, which is just a list of DDLIB Word objects
sent = []
for i,t in enumerate(tokens):
sent.append(ddlib.Word(
begin_char_offset=None,
end_char_offset=None,
word=t,
lemma=lemmas[i],
pos=pos_tags[i],
ner=ner_tags[i],
dep_par=dep_parents[i] - 1, # Note that as stored from CoreNLP 0 is ROOT, but for DDLIB -1 is ROOT
dep_label=dep_types[i]))
# Create DDLIB Spans for the two person mentions
p1_span = ddlib.Span(begin_word_id=p1_begin_index, length=(p1_end_index-p1_begin_index+1))
p2_span = ddlib.Span(begin_word_id=p2_begin_index, length=(p2_end_index-p2_begin_index+1))
# Generate the generic features using DDLIB
for feature in ddlib.get_generic_features_relation(sent, p1_span, p2_span):
yield [p1_id, p2_id, feature]
Note that getting the input for this UDF requires joining the person_mention and sentences tables:
function extract_spouse_features over (
p1_id text,
p2_id text,
p1_begin_index int,
p1_end_index int,
p2_begin_index int,
p2_end_index int,
doc_id text,
sent_index int,
tokens text[],
lemmas text[],
pos_tags text[],
ner_tags text[],
dep_types text[],
dep_tokens int[]
) returns rows like spouse_feature
implementation "udf/extract_spouse_features.py" handles tsv lines.
spouse_feature += extract_spouse_features(
p1_id, p2_id, p1_begin_index, p1_end_index, p2_begin_index, p2_end_index,
doc_id, sent_index, tokens, lemmas, pos_tags, ner_tags, dep_types, dep_tokens
) :-
person_mention(p1_id, _, doc_id, sent_index, p1_begin_index, p1_end_index),
person_mention(p2_id, _, doc_id, sent_index, p2_begin_index, p2_end_index),
sentences(doc_id, sent_index, _, tokens, lemmas, pos_tags, ner_tags, _, dep_types, dep_tokens).
Again, to run, just compile and execute as in previous steps.
deepdive compile && deepdive do spouse_feature
If we take a look at a sample of the extracted features, they will look roughly like the following:
deepdive query '| 20 ?- spouse_feature(_, _, feature).'
feature
-------------------------------------------------------
WORD_SEQ_[accepted a plea deal and testified against]
LEMMA_SEQ_[accept a plea deal and testify against]
NER_SEQ_[O O O O O O O]
POS_SEQ_[VBD DT NN NN CC VBD IN]
W_LEMMA_L_1_R_1_[.]_[.]
W_NER_L_1_R_1_[O]_[O]
W_LEMMA_L_2_R_1_[Gissendaner .]_[.]
W_NER_L_2_R_1_[PERSON O]_[O]
W_LEMMA_L_3_R_1_[against Gissendaner .]_[.]
W_NER_L_3_R_1_[O PERSON O]_[O]
NGRAM_1_[accept]
NGRAM_2_[accept a]
NGRAM_3_[accept a plea]
NGRAM_1_[a]
NGRAM_2_[a plea]
NGRAM_3_[a plea deal]
NGRAM_1_[plea]
NGRAM_2_[plea deal]
NGRAM_3_[plea deal and]
NGRAM_1_[deal]
(20 rows)
Now we have generated what looks more like the standard input to a machine learning problem—a set of objects, represented by sets of features, which we want to classify (here, as true or false mentions of a spousal relation). However, we don't have any supervised labels (i.e., a set of correct answers) for a machine learning algorithm to learn from! In most real world applications, a sufficiently large set of supervised labels is not available. With DeepDive, we take the approach sometimes referred to as distant supervision or data programming, where we instead generate a noisy set of labels using a mix of mappings from secondary datasets and other heuristic rules.
2. Distant supervision with data and rules
In this section, we'll use distant supervision (or 'data programming') to provide a noisy set of labels for candidate relation mentions, with which we will train a machine learning model.
We'll describe two basic categories of approaches:
- Mapping from secondary data for distant supervision
- Using heuristic rules for distant supervision
Finally, we'll describe a simple majority-vote approach to resolving multiple labels per example, which can be implemented within DDlog.
2.1. Mapping from secondary data for distant supervision
First, we'll try using an external structured dataset of known married couples, from DBpedia, to distantly supervise our dataset. We'll download the relevant data, and then map it to our candidate spouse relations.
Extracting and downloading the DBpedia data
Our goal is to first extract a collection of known married couples from DBpedia and then load this into the spouses_dbpedia table in our database.
To extract known married couples, we use the DBpedia dump present in Google's BigQuery platform.
First we extract the URI, name and spouse information from the DBpedia person table records in BigQuery for which the field name is not NULL.
We use the following query:
SELECT URI,name, spouse
FROM [fh-bigquery:dbpedia.person]
where name <> "NULL"
We store the result of the above query in a local project table dbpedia.validnames and perform a self-join to obtain the pairs of married couples.
SELECT t1.name, t2.name
FROM [dbpedia.validnames] AS t1
JOIN EACH [dbpedia.validnames] AS t2
ON t1.spouse = t2.URI
The output of the above query is stored in a new table named dbpedia.spouseraw.
Finally, we use the following query to remove symmetric duplicates.
SELECT p1, p2
FROM (SELECT t1_name as p1, t2_name as p2 FROM [dbpedia.spouseraw]),
(SELECT t2_name as p1, t1_name as p2 FROM [dbpedia.spouseraw])
WHERE p1 < p2
The output of this query is stored in a local file.
The file contains duplicate rows (BigQuery does not support distinct).
It also contains noisy rows where the name field contains a string where the given name family name and multiple aliases were concatenated and reported in a string including the characters { and }.
Using the Unix commands sed, sort and uniq we first remove the lines containing characters { and } and then duplicate entries.
This results in an input file spouses_dbpedia.csv containing 6,126 entries of married couples.
Loading DBpedia data to database
We compress and store spouses_dbpedia.csv under the path:
input/spouses_dbpedia.csv.bz2
Notice that for DeepDive to load the data to the corresponding database table, the name of the input data again has to be stored in the directory input/ and has the same name as the target database table.
To load the data we execute the command:
deepdive do spouses_dbpedia
Now the database should include tuples that look like the following:
deepdive query '| 20 ?- spouses_dbpedia(name1, name2).'
name1 | name2
------------------------------------+--------------------------------------------------------
A. A. Gill | Amber Rudd
Aamir Ali Malik | Sanjeeda Shaikh
Abimael Guzmán | Augusta la Torre
Abraham Jacobi | Mary Corinna Putnam Jacobi
Addison Adrienne Forbes Montgomery | Derek Christopher Shepherd
Alan Crosland | Elaine Hammerstein
Albert | Anna Marie of Brunswick-Lüneburg
Albert | Archduchess Margarethe Klementine of Austria
Albert | Dorothea
Albert | Isabella Clara Eugenia
Albert | Karoline Friederike Franziska Stephanie Amalie Cecilie
Albert | Richardis of Schwerin
Aleksander Ludwik Radziwiłł | Katarzyna Eugenia Tyszkiewicz
Alessia Merz | Fabio Bazzani
Alexis Denisof | Alyson Hannigan
Alfonso XII | Maria Christina of Austria
Alice of Namur | Baldwin IV Count of Hainaut
Amine Gemayel | Joyce Gemayel
Andrés Pastrana Arango | Nohra Puyana Bickenbach
Andrew Cymek | Brigitte Kingsley
(20 rows)
Supervising spouse candidates with DBpedia data
First we'll declare a new table where we'll store the labels (referring to the spouse candidate mentions), with an integer value (True=1, False=-1) and a description (rule_id):
spouse_label(
p1_id text,
p2_id text,
label int,
rule_id text
).
Next we'll implement a simple distant supervision rule which labels any spouse mention candidate with a pair of names appearing in DBpedia as true:
# distant supervision using data from DBpedia
spouse_label(p1,p2, 1, "from_dbpedia") :-
spouse_candidate(p1, p1_name, p2, p2_name), spouses_dbpedia(n1, n2),
[ lower(n1) = lower(p1_name), lower(n2) = lower(p2_name) ;
lower(n2) = lower(p1_name), lower(n1) = lower(p2_name) ].
It should be noted that there are many clear ways in which this rule could be improved (fuzzy matching, more restrictive conditions, etc.), but this serves as an example of one major type of distant supervision rule.
2.2. Using heuristic rules for distant supervision
We can also create a supervision rule which does not rely on any secondary structured dataset like DBpedia, but instead just uses some heuristic.
We set up a DDlog function, supervise, which uses a UDF containing several heuristic rules over the mention and sentence attributes:
function supervise over (
p1_id text, p1_begin int, p1_end int,
p2_id text, p2_begin int, p2_end int,
doc_id text,
sentence_index int,
sentence_text text,
tokens text[],
lemmas text[],
pos_tags text[],
ner_tags text[],
dep_types text[],
dep_tokens int[]
) returns (
p1_id text, p2_id text, label int, rule_id text
)
implementation "udf/supervise_spouse.py" handles tsv lines.
spouse_label += supervise(
p1_id, p1_begin, p1_end,
p2_id, p2_begin, p2_end,
doc_id, sentence_index, sentence_text,
tokens, lemmas, pos_tags, ner_tags, dep_types, dep_token_indexes
) :-
spouse_candidate(p1_id, _, p2_id, _),
person_mention(p1_id, p1_text, doc_id, sentence_index, p1_begin, p1_end),
person_mention(p2_id, p2_text, _, _, p2_begin, p2_end),
sentences(
doc_id, sentence_index, sentence_text,
tokens, lemmas, pos_tags, ner_tags, _, dep_types, dep_token_indexes
).
The Python UDF named udf/supervise_spouse.py contains several heuristic rules:
- Candidates with person mentions that are too far apart in the sentence are marked as false.
- Candidates with person mentions that have another person in between are marked as false.
- Candidates with person mentions that have words like "wife" or "husband" in between are marked as true.
- Candidates with person mentions that have "and" in between and "married" after are marked as true.
- Candidates with person mentions that have familial relation words in between are marked as false.
#!/usr/bin/env python
from deepdive import *
import random
from collections import namedtuple
SpouseLabel = namedtuple('SpouseLabel', 'p1_id, p2_id, label, type')
@tsj_extractor
@returns(lambda
p1_id = "text",
p2_id = "text",
label = "int",
rule_id = "text",
:[])
# heuristic rules for finding positive/negative examples of spouse relationship mentions
def supervise(
p1_id="text", p1_begin="int", p1_end="int",
p2_id="text", p2_begin="int", p2_end="int",
doc_id="text", sentence_index="int",
tokens="text[]", lemmas="text[]", pos_tags="text[]", ner_tags="text[]",
dep_types="text[]", dep_token_indexes="int[]",
):
# Constants
MARRIED = frozenset(["wife", "husband"])
FAMILY = frozenset(["mother", "father", "sister", "brother", "brother-in-law"])
MAX_DIST = 10
# Common data objects
p1_end_idx = min(p1_end, p2_end)
p2_start_idx = max(p1_begin, p2_begin)
p2_end_idx = max(p1_end,p2_end)
intermediate_lemmas = lemmas[p1_end_idx+1:p2_start_idx]
intermediate_ner_tags = ner_tags[p1_end_idx+1:p2_start_idx]
tail_lemmas = lemmas[p2_end_idx+1:]
spouse = SpouseLabel(p1_id=p1_id, p2_id=p2_id, label=None, type=None)
# Rule: Candidates that are too far apart
if len(intermediate_lemmas) > MAX_DIST:
yield spouse._replace(label=-1, type='neg:far_apart')
# Rule: Candidates that have a third person in between
if 'PERSON' in intermediate_ner_tags:
yield spouse._replace(label=-1, type='neg:third_person_between')
# Rule: Sentences that contain wife/husband in between
# (<P1>)([ A-Za-z]+)(wife|husband)([ A-Za-z]+)(<P2>)
if len(MARRIED.intersection(intermediate_lemmas)) > 0:
yield spouse._replace(label=1, type='pos:wife_husband_between')
# Rule: Sentences that contain and ... married
# (<P1>)(and)?(<P2>)([ A-Za-z]+)(married)
if ("and" in intermediate_lemmas) and ("married" in tail_lemmas):
yield spouse._replace(label=1, type='pos:married_after')
# Rule: Sentences that contain familial relations:
# (<P1>)([ A-Za-z]+)(brother|stster|father|mother)([ A-Za-z]+)(<P2>)
if len(FAMILY.intersection(intermediate_lemmas)) > 0:
yield spouse._replace(label=-1, type='neg:familial_between')
Note that the rough theory behind this approach is that we don't need high-quality (e.g., hand-labeled) supervision to learn a high quality model. Instead, using statistical learning, we can in fact recover high-quality models from a large set of low-quality or noisy labels.
2.3. Resolving multiple labels per example with majority vote
Finally, we implement a very simple majority vote procedure, all in DDlog, for resolving scenarios where a single spouse candidate mention has multiple conflicting labels. First, we sum the labels (which are all -1, 0, or 1):
spouse_label_resolved(p1_id, p2_id, SUM(vote)) :- spouse_label(p1_id, p2_id, vote, rule_id).
Then, we simply threshold and add these labels to our decision variable table has_spouse (see next section for details here):
has_spouse(p1_id, p2_id) = if l > 0 then TRUE
else if l < 0 then FALSE
else NULL end :- spouse_label_resolved(p1_id, p2_id, l).
We additionally make sure that all spouse candidate mentions not labeled by a rule are also included in this table:
has_spouse(p1, p2) = NULL :- spouse_candidate(p1, _, p2, _).
Once again, to execute all of the above, just run the following command:
deepdive compile && deepdive do has_spouse
Recall that deepdive do will execute all upstream tasks as well, so this will execute all of the previous steps!
Now, we can take a brief look at how many candidates are supervised by different rules, which will look something like the table below. Obviously, the counts will vary depending on your input corpus.
deepdive query 'rule, @order_by COUNT(1) ?- spouse_label(p1,p2, label, rule).'
rule | COUNT(1)
--------------------------+----------
pos:married_after | 556
from_dbpedia | 1010
neg:familial_between | 24944
pos:wife_husband_between | 53032
neg:third_person_between | 194898
neg:far_apart | 305488
| 731982
(7 rows)
3. Learning and inference: model specification
Now, we need to specify the actual model that DeepDive will perform learning and inference over. At a high level, this boils down to specifying three things:
What are the variables of interest that we want DeepDive to predict for us?
What are the features for each of these variables?
What are the connections between the variables?
One we have specified the model in this way, DeepDive will learn the parameters of the model (the weights of the features and potentially the connections between variables), and then perform statistical inference over the learned model to determine the probability that each variable of interest is true.
For more advanced users: we are specifying a factor graph where the features are unary factors, and then using SGD and Gibbs sampling for learning and inference. Further technical detail is available here.
3.1. Specifying prediction variables
In our case, we have one variable to predict per spouse candidate mention, namely, is this mention actually indicating a spousal relation or not?
In other words, we want DeepDive to predict the value of a Boolean variable for each spouse candidate mention, indicating whether it is true or not.
We specify this in app.ddlog as follows:
has_spouse?(
p1_id text,
p2_id text
).
DeepDive will predict not only the value of these variables, but also the marginal probabilities, i.e., the confidence level that DeepDive has for each individual prediction.
3.2. Specifying features
Next, we indicate (i) that each has_spouse variable will be connected to the features of the corresponding spouse_candidate row, (ii) that we wish DeepDive to learn the weights of these features from our distantly supervised data, and (iii) that the weight of a specific feature across all instances should be the same, as follows:
@weight(f)
has_spouse(p1_id, p2_id) :-
spouse_candidate(p1_id, _, p2_id, _),
spouse_feature(p1_id, p2_id, f).
3.3. Specifying connections between variables
Finally, we can specify dependencies between the prediction variables, with either learned or given weights.
Here, we'll specify two such rules, with fixed (given) weights that we specify.
First, we define a symmetry connection, namely specifying that if the model thinks a person mention p1 and a person mention p2 indicate a spousal relationship in a sentence, then it should also think that the reverse is true, i.e., that p2 and p1 indicate one too:
@weight(3.0)
has_spouse(p1_id, p2_id) => has_spouse(p2_id, p1_id) :-
spouse_candidate(p1_id, _, p2_id, _).
Next, we specify a rule that the model should be strongly biased towards finding one marriage indication per person mention. We do this inversely, using a negative weight, as follows:
@weight(-1.0)
has_spouse(p1_id, p2_id) => has_spouse(p1_id, p3_id) :-
spouse_candidate(p1_id, _, p2_id, _),
spouse_candidate(p1_id, _, p3_id, _).
3.4. Performing learning and inference
Finally, to perform learning and inference using the specified model, we need to run the following command:
deepdive compile && deepdive do probabilities
This will ground the model based on the data in the database, learn the weights, infer the expectations or marginal probabilities of the variables in the model, and then load them back to the database.
Let's take a look at the probabilities inferred by DeepDive for the has_spouse variables.
deepdive sql "SELECT p1_id, p2_id, expectation FROM has_spouse_inference ORDER BY random() LIMIT 20"
p1_id | p2_id | expectation
-----------------------------------------------+-----------------------------------------------+-------------
1d1cff32-f332-41c3-9c18-57f44b5d7b02_4_12_12 | 1d1cff32-f332-41c3-9c18-57f44b5d7b02_4_6_7 | 0.171
5c467615-1dfc-4399-be91-6979cf6eade8_16_20_20 | 5c467615-1dfc-4399-be91-6979cf6eade8_16_22_22 | 0.952
bdecf4c6-81eb-4851-b30c-84f80ff58048_10_20_20 | bdecf4c6-81eb-4851-b30c-84f80ff58048_10_26_26 | 0.001
fd2c5043-52ac-44e6-af50-9d3c32f6b4ce_49_24_25 | fd2c5043-52ac-44e6-af50-9d3c32f6b4ce_49_29_30 | 0.034
f55c8585-893f-4e06-b904-9a1af91586c2_17_6_7 | f55c8585-893f-4e06-b904-9a1af91586c2_17_2_3 | 0.107
b9e56701-b378-4049-af23-5473b4878174_8_39_40 | b9e56701-b378-4049-af23-5473b4878174_8_13_13 | 0
17166b09-c02b-436e-82ec-bfd5afc5f23d_11_7_8 | 17166b09-c02b-436e-82ec-bfd5afc5f23d_11_5_5 | 0.163
15aa10bc-cd54-4956-9fb8-a61a850b86e3_50_17_18 | 15aa10bc-cd54-4956-9fb8-a61a850b86e3_50_9_15 | 0.993
c0e7a274-dac0-4913-96cc-2ec5209c52e0_4_1_1 | c0e7a274-dac0-4913-96cc-2ec5209c52e0_4_40_40 | 0
15ea7ce9-b341-40b1-a0a9-b784ebd010fc_7_12_12 | 15ea7ce9-b341-40b1-a0a9-b784ebd010fc_7_5_5 | 0.04
f7ab24b8-ec6d-4293-9022-648818b7cbfe_17_9_9 | f7ab24b8-ec6d-4293-9022-648818b7cbfe_17_1_1 | 0.017
b0430512-20d6-4c62-aa06-988a3bdaa1c3_30_37_37 | b0430512-20d6-4c62-aa06-988a3bdaa1c3_30_27_27 | 0.018
147396b5-a728-4fc7-989b-5479c617d484_32_16_17 | 147396b5-a728-4fc7-989b-5479c617d484_32_5_6 | 0
72dff05a-4092-4724-b1c5-28e47d5626ce_17_5_5 | 72dff05a-4092-4724-b1c5-28e47d5626ce_17_3_3 | 0.992
c502650f-d0ca-4298-ab96-64ceebf70626_17_10_10 | c502650f-d0ca-4298-ab96-64ceebf70626_17_8_8 | 0.533
a8391c1e-12d1-4178-9d79-cf69c94d58f8_13_1_1 | a8391c1e-12d1-4178-9d79-cf69c94d58f8_13_5_5 | 0.038
7e5b8b2b-623d-423d-9da9-8ecb1f3a8c26_20_11_12 | 7e5b8b2b-623d-423d-9da9-8ecb1f3a8c26_20_0_1 | 0
954b6e8e-dce8-47e5-a769-c058ee6dbc00_2_15_15 | 954b6e8e-dce8-47e5-a769-c058ee6dbc00_2_12_12 | 0.059
71f4101b-42c5-4536-bee9-cffdd840da74_34_0_0 | 71f4101b-42c5-4536-bee9-cffdd840da74_34_2_2 | 0.978
09e8cfe7-06e9-4abd-8269-db8a90192552_50_0_0 | 09e8cfe7-06e9-4abd-8269-db8a90192552_50_10_11 | 0.002
(20 rows)
4. Error analysis and debugging
After finishing a pass of writing and running the DeepDive application, the first thing we want to see is how good the results are. In this section, we describe how DeepDive's interactive tools can be used for viewing the results as well as error analysis and debugging.
4.1. Calibration Plots
DeepDive provides calibration plots to see how well the expectations computed by the system are calibrated.
The following command generates a plot for each variable under run/model/calibration-plots/.
deepdive do calibration-plots
It will produce a file run/model/calibration-plots/has_spouse.png that holds three plots as shown below:
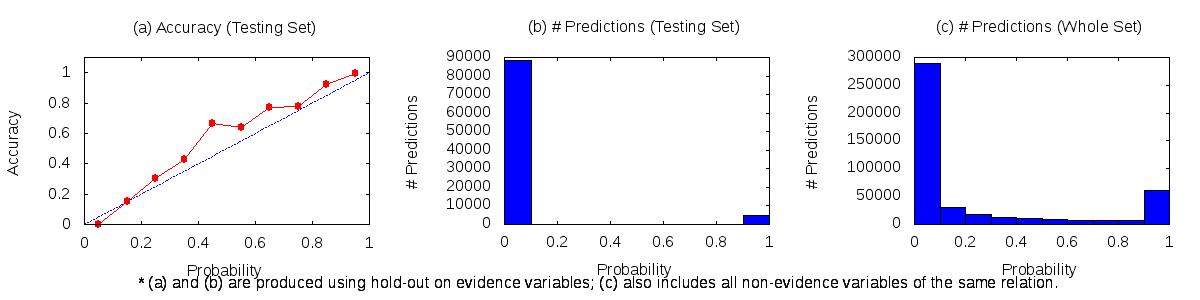
Refer to the full documentation on calibration data for more detail on how to interpret the plots and take actions.
4.2. Browsing data with Mindbender
Mindbender is the name of the tool that provides an interactive user interface to DeepDive. It can be used for browsing any data that has been loaded into DeepDive and produced by it.
Browsing input corpus
We need to give hints to DeepDive about which part of the data we want to browse using DDlog's annotation.
For example, on the articles relation we declared earlier in app.ddlog, we can sprinkle some annotations such as @source, @key, and @searchable, as the following.
@source
articles(
@key
id text,
@searchable
content text
).
Next, if we run the following command, DeepDive will create and populate a search index according to these hints.
mindbender search update
To access the populated search index through a web browser, run:
mindbender search gui
Then, point your browser to the URL that appears after the command (typically http://localhost:8000) to see a view that looks like the following:
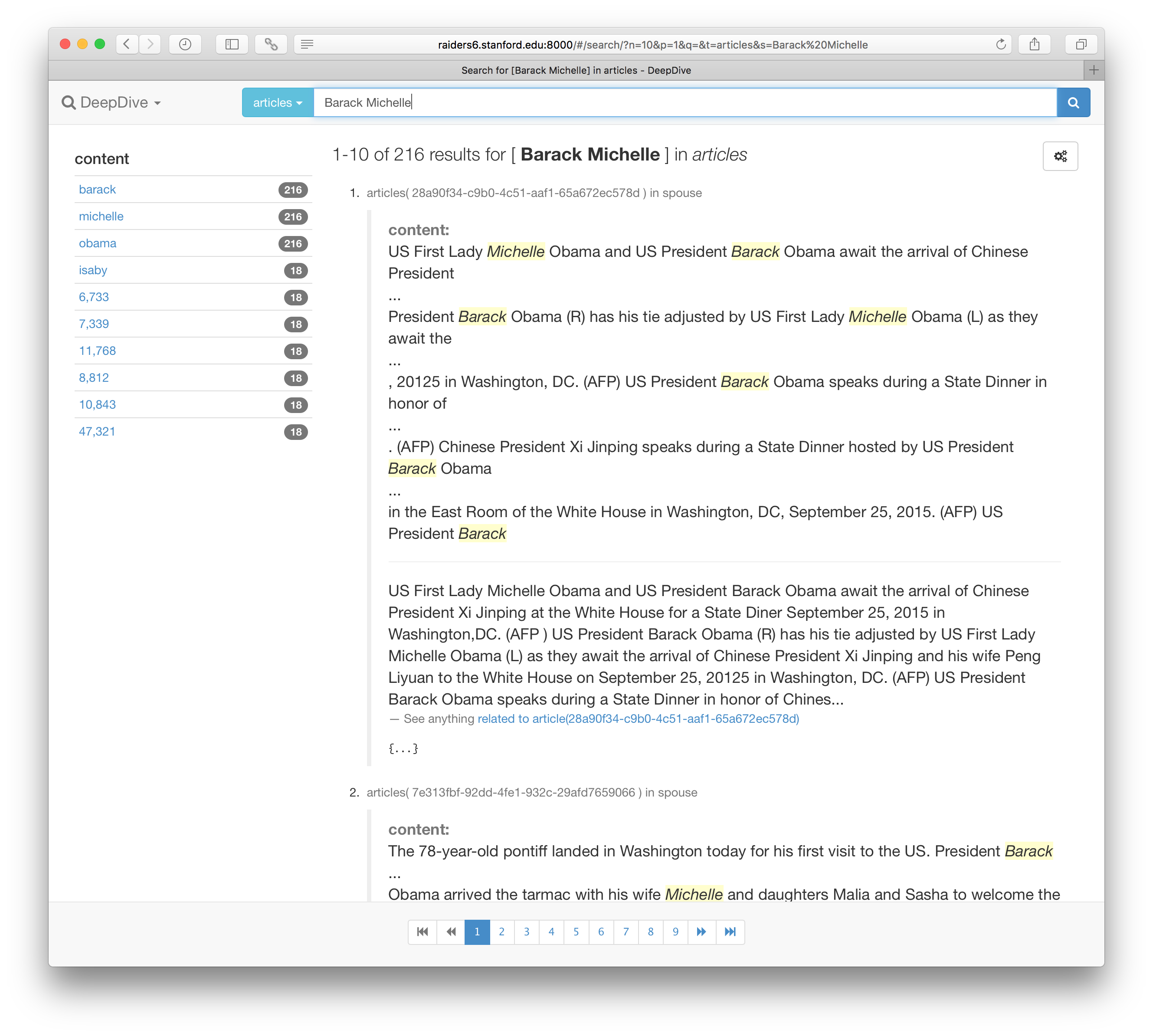
Browsing result data
To browse the results, we can add annotations to the inferred relations and how they relate to their source relations.
For example, the @extraction and @references annotations in the following DDlog declaration tells DeepDive that the variable relation has_spouse is inferred from pairs of person_mention.
@extraction
has_spouse?(
@key
@references(relation="person_mention", column="mention_id", alias="p1")
p1_id text,
@key
@references(relation="person_mention", column="mention_id", alias="p2")
p2_id text
).
The relation person_mention as well as the relations it references should have similar annotations (see the complete app.ddlog code for full detail).
Then, repeating the commands to update the search index and load the user interface will allow us to browse the expected marginal probabilities of has_spouse as well.
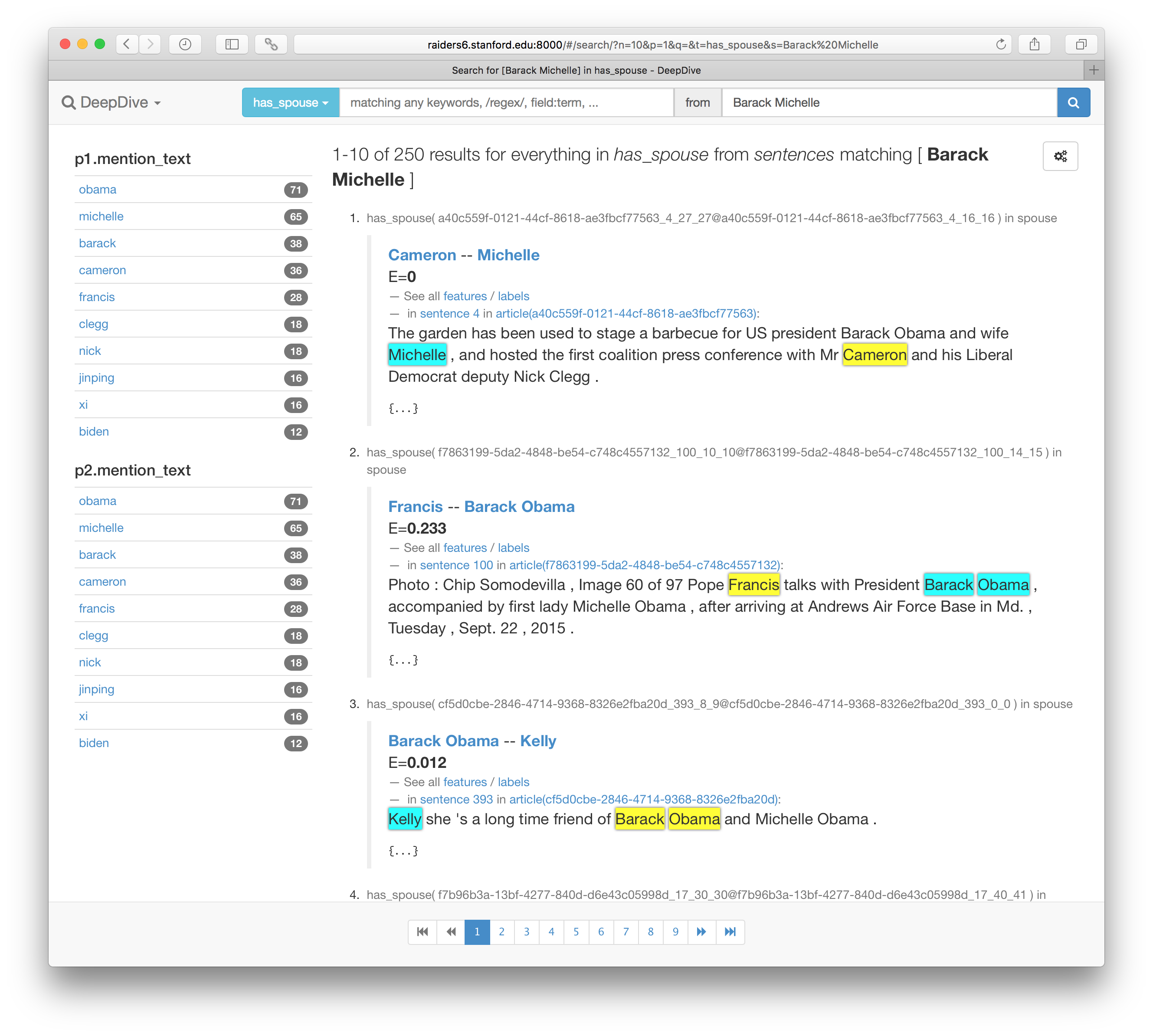
Customizing how data is presented
In fact, the screenshots above are showing the data presented using a carefully prepared set of templates under mindbender/search-templates/.
In these AngularJS templates, virtually anything you can program in HTML/CSS/JavaScript/CoffeeScript can be added to present the data that is ideal for human consumption (e.g., highlighted text spans rather than token indexes).
Please see the documentation about customizing the presentation for further detail.
4.3. Estimating precision with Mindtagger
Mindtagger, which is part of the Mindbender tool suite, assists data labeling tasks to quickly assess the precision and/or recall of the extraction.
We show how Mindtagger helps us perform a labeling task to estimate the precision of the extraction.
The necessary set of files shown below already exist in the example under labeling/has_spouse-precision/.
Preparing a data labeling task
First, we can take a random sample of 100 examples from has_spouse relation whose expectation is higher than or equal to a 0.9 threshold as shown in the following SQL query, and store them in a file called has_spouse.csv.
deepdive sql eval "
SELECT hsi.p1_id
, hsi.p2_id
, s.doc_id
, s.sentence_index
, hsi.label
, hsi.expectation
, s.tokens
, pm1.mention_text AS p1_text
, pm1.begin_index AS p1_start
, pm1.end_index AS p1_end
, pm2.mention_text AS p2_text
, pm2.begin_index AS p2_start
, pm2.end_index AS p2_end
FROM has_spouse_inference hsi
, person_mention pm1
, person_mention pm2
, sentences s
WHERE hsi.p1_id = pm1.mention_id
AND pm1.doc_id = s.doc_id
AND pm1.sentence_index = s.sentence_index
AND hsi.p2_id = pm2.mention_id
AND pm2.doc_id = s.doc_id
AND pm2.sentence_index = s.sentence_index
AND expectation >= 0.9
ORDER BY random()
LIMIT 100
" format=csv header=1 >labeling/has_spouse-precision/has_spouse.csv
We also prepare the mindtagger.conf and template.html files under labeling/has_spouse-precision/ that look like the following:
title: Labeling task for estimating has_spouse precision
items: {
file: has_spouse.csv
key_columns: [p1_id, p2_id]
}
template: template.html
<mindtagger mode="precision">
<template for="each-item">
<strong title="item_id: "> -- </strong>
with expectation <strong></strong> appeared in:
<blockquote>
<big mindtagger-word-array="item.tokens" array-format="json">
<mindtagger-highlight-words from="item.p1_start" to="item.p1_end" with-style="background-color: yellow;"/>
<mindtagger-highlight-words from="item.p2_start" to="item.p2_end" with-style="background-color: cyan;"/>
</big>
</blockquote>
<div>
<div mindtagger-item-details></div>
</div>
</template>
<template for="tags">
<span mindtagger-adhoc-tags></span>
<span mindtagger-note-tags></span>
</template>
</mindtagger>
Labeling data with Mindtagger
Mindtagger can then be started for the task using the following command:
mindbender tagger labeling/has_spouse-precision/mindtagger.conf
Then, point your browser to the URL that appears after the command (typically http://localhost:8000) to see a dedicated user interface for labeling data that looks like the following:
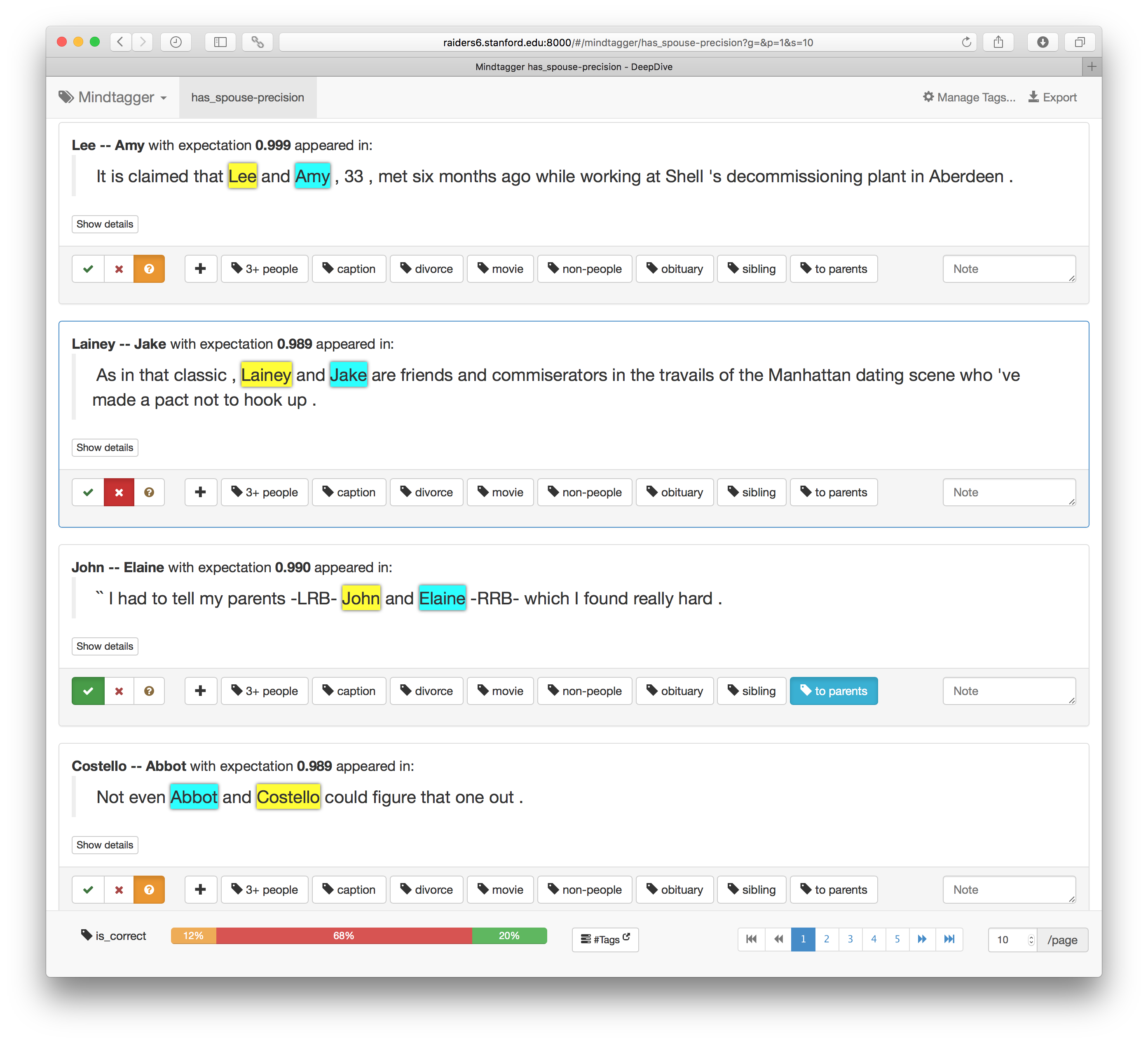
We can quickly label the sampled 100 examples using the intuitive user interface with buttons for correct/incorrect tags. It also supports keyboard shortcuts for entering labels and moving between items. (Press the ? key to view all supported keys.) How many were labeled correct, as well as other tags, are shown in the "Tags" dropdown at the top right corner as shown below.
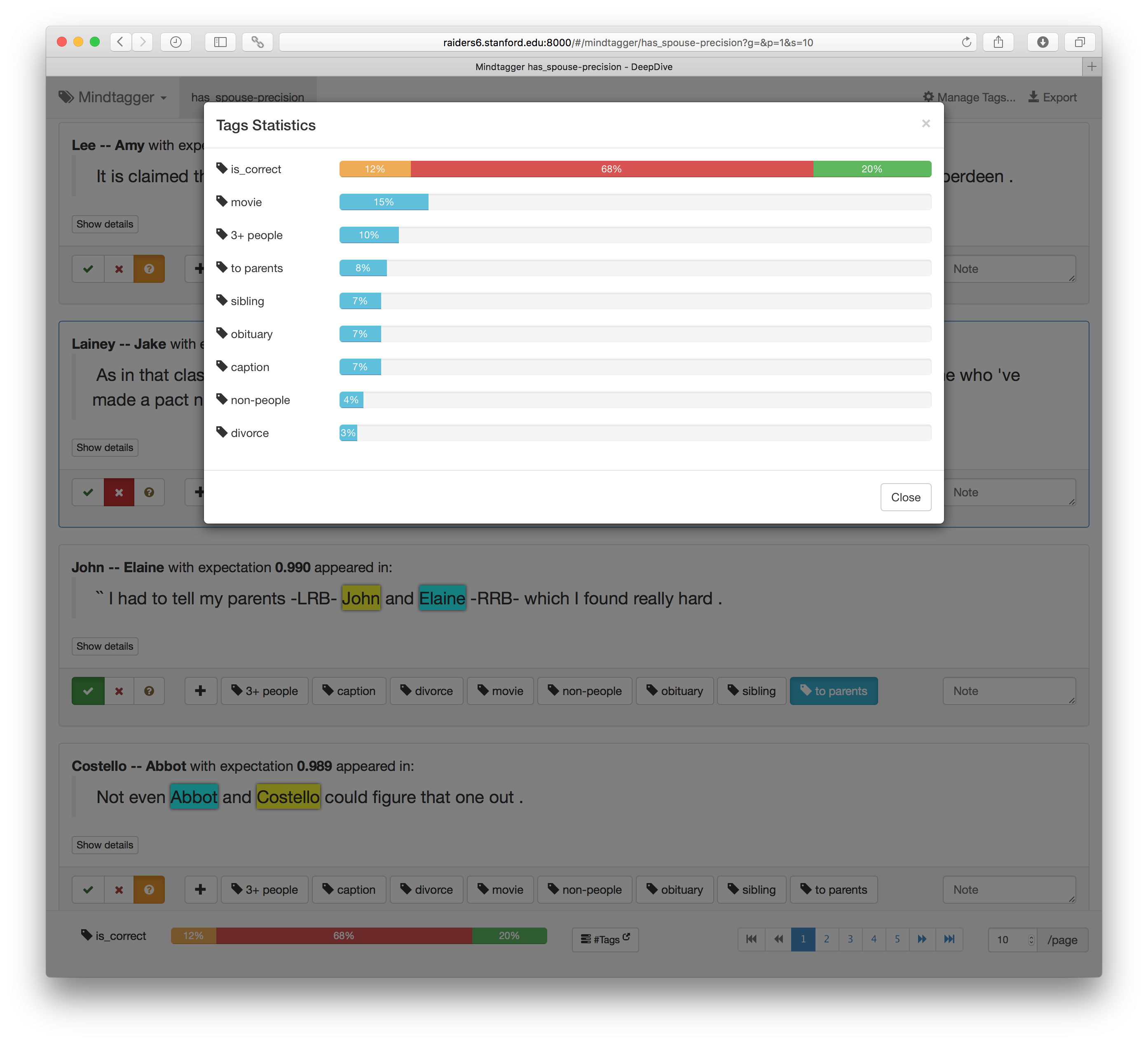
The collected tags can also be exported in various format for post-processing.
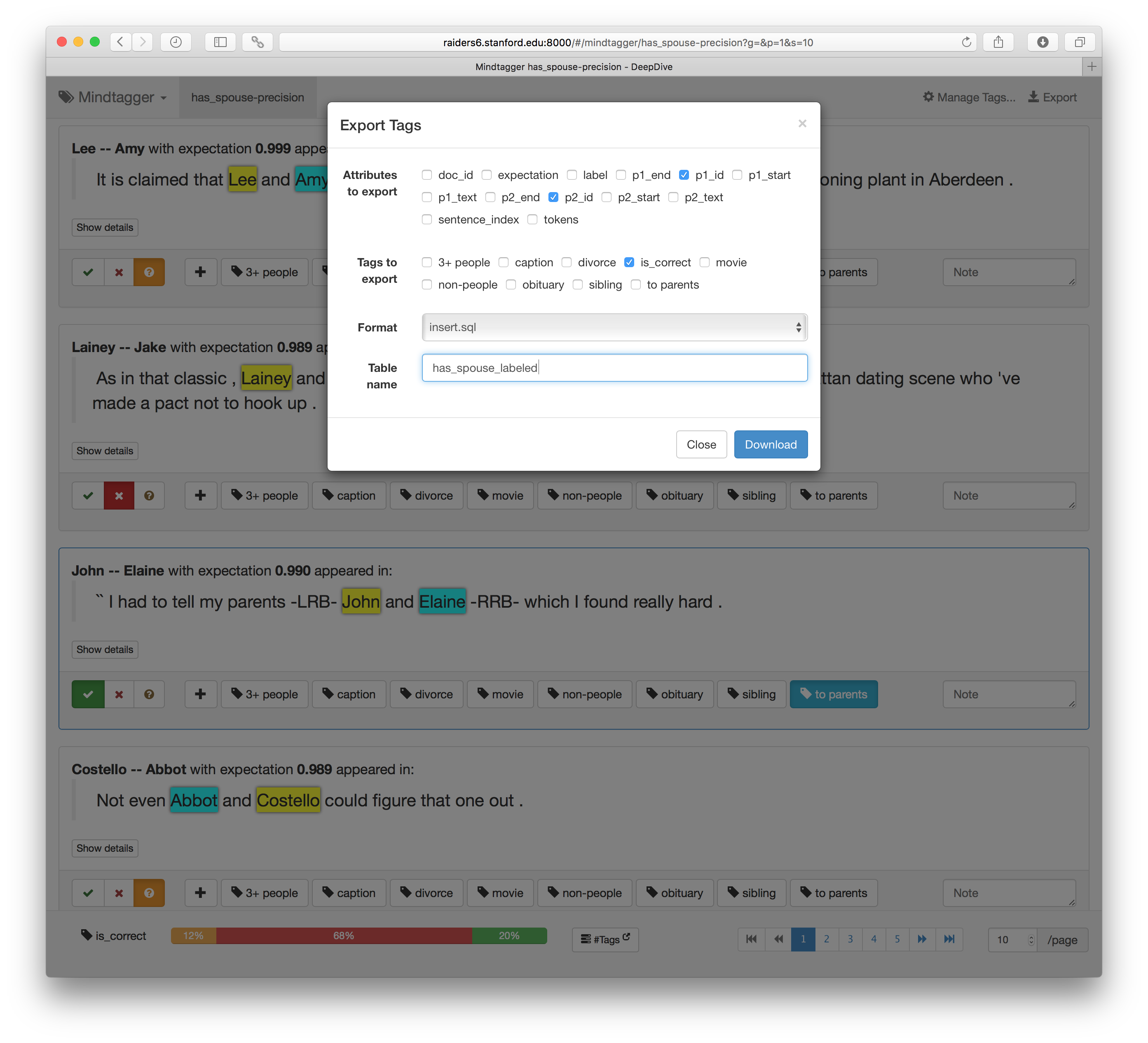
For further detail, see the documentation about labeling data.
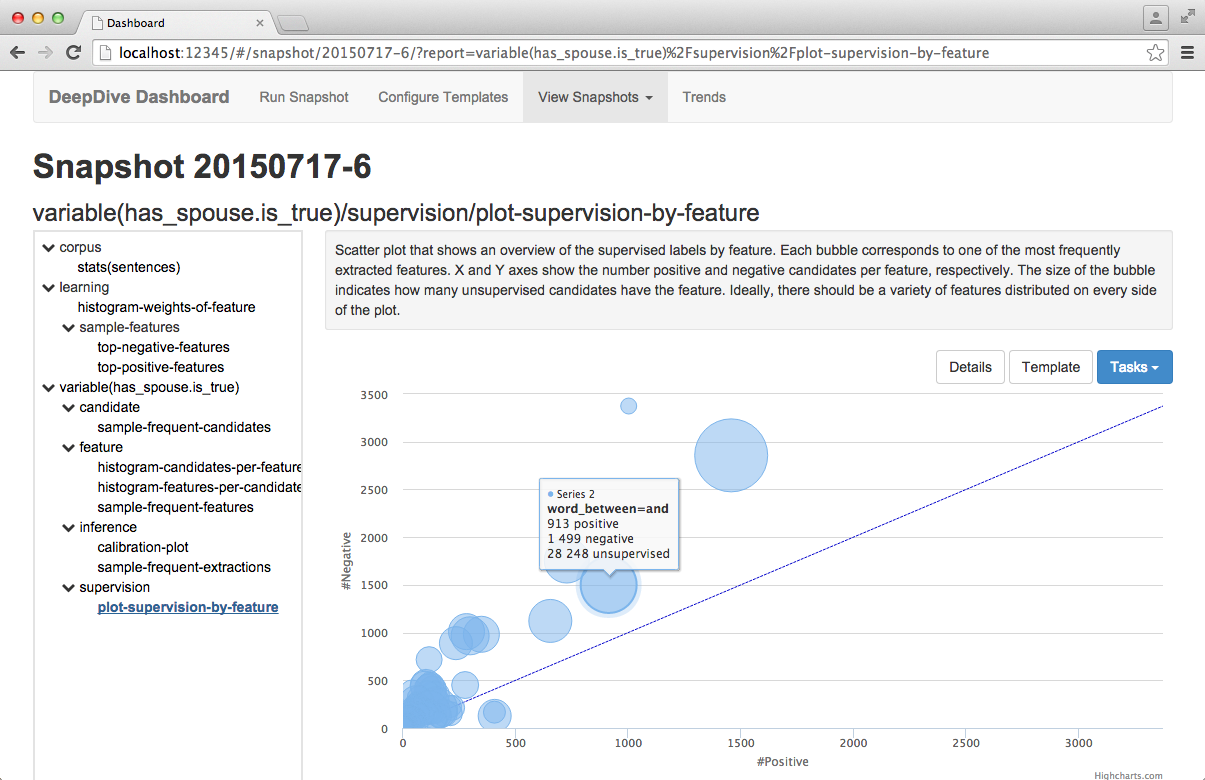
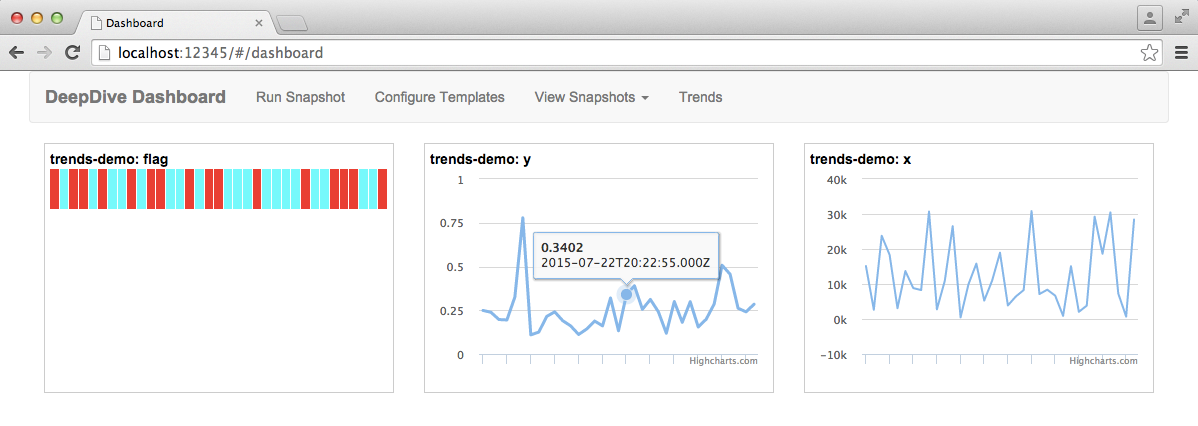
4.4. Monitoring statistics with Dashboard
Dashboard provides a way to monitor various descriptive statistics of the data products after each pass of DeepDive improvements. We can use a combination of SQL, any Bash script, and Markdown in each report template that produces a report, and we can produce a collection of them as a snapshot against the data extracted by DeepDive. Dashboard provides a structure to manage those templates and instantiate them in a sophisticated way using parameters. It provides a graphical interface for visualizing the collected statistics and trends as shown below. Refer to the full documentation on Dashboard to set up your own set of reports.